 Back
BackGene Expression: From DNA to Protein (Central Dogma, Transcription, Translation, and Mutations)
Study Guide - Smart Notes

Gene Expression: The Central Dogma
Overview of the Central Dogma
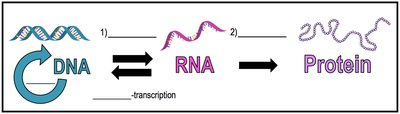
The central dogma of molecular biology describes the unidirectional flow of genetic information within a biological system: from DNA to RNA to protein. This process involves two main steps: transcription (DNA to RNA) and translation (RNA to protein). The central dogma is fundamental to understanding how genotypes are expressed as phenotypes.
Transcription: The process of synthesizing RNA from a DNA template.
Translation: The process of synthesizing proteins using the information encoded in mRNA.
Gene Expression: The full process by which genetic information is used to synthesize gene products (RNA or protein).
Transcription: From DNA to RNA
Introduction to Transcription
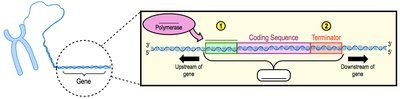
Transcription is the process by which a segment of DNA (a gene) is used as a template to synthesize a complementary RNA molecule. Genes are specific sequences of DNA that encode functional products, typically proteins.
Promoter: A DNA sequence where transcription begins; site of RNA polymerase attachment.
Terminator: A DNA sequence where transcription ends.
RNA Polymerase: The enzyme that synthesizes RNA from the DNA template, without the need for a primer.
Upstream/Downstream: Terms describing directionality relative to the transcription start site.
DNA Strands in Transcription
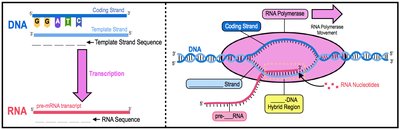
DNA consists of two strands: the coding strand (sense) and the template strand (antisense). The RNA sequence produced during transcription is complementary to the template strand and nearly identical to the coding strand, except that RNA contains uracil (U) instead of thymine (T).
Base Pairing: A pairs with U (in RNA), C pairs with G.
Directionality: RNA is synthesized in the 5' to 3' direction.
Steps of Transcription
Transcription occurs in three main steps: initiation, elongation, and termination.
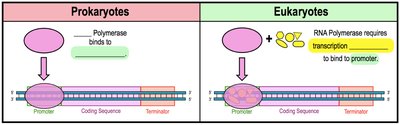
Initiation: RNA polymerase binds to the promoter and unwinds the DNA.
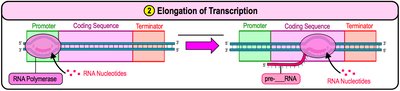
Elongation: RNA polymerase moves along the DNA, synthesizing RNA by adding nucleotides complementary to the template strand.
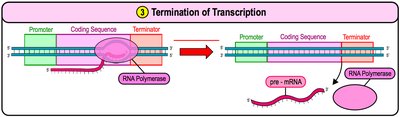
Termination: RNA polymerase reaches the terminator sequence and releases the newly synthesized RNA.
RNA Processing in Eukaryotes
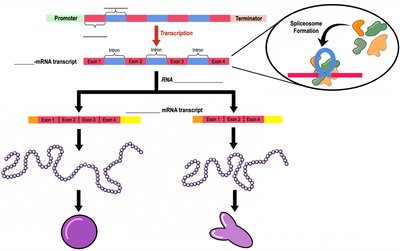
RNA Processing and Splicing
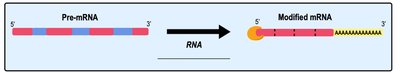
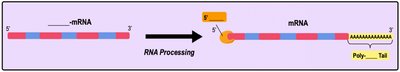
In eukaryotes, the initial RNA transcript (pre-mRNA) undergoes several modifications before becoming mature mRNA ready for translation. These modifications include the addition of a 5' cap, a poly-A tail, and the removal of non-coding sequences (introns) through splicing.
5' Cap: Modified guanine nucleotide added to the 5' end for stability and ribosome binding.
Poly-A Tail: String of adenine nucleotides added to the 3' end for stability and export from the nucleus.
Splicing: Removal of introns (non-coding regions) and joining of exons (coding regions) by the spliceosome.
Alternative Splicing: Allows a single gene to code for multiple proteins by varying the combination of exons included in the final mRNA.
Types of RNA
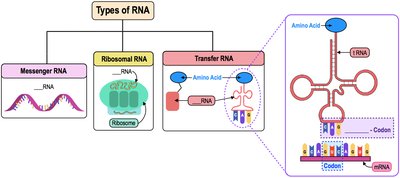
Major Types of RNA and Their Functions
Cells use several types of RNA, each with a distinct function in gene expression:
Messenger RNA (mRNA): Carries genetic information from DNA to the ribosome for protein synthesis. Contains codons (three-nucleotide sequences) that specify amino acids.
Ribosomal RNA (rRNA): Structural and catalytic component of ribosomes.
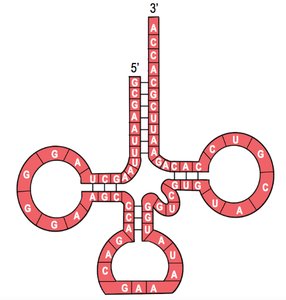
Transfer RNA (tRNA): Brings amino acids to the ribosome during translation. Contains anticodons complementary to mRNA codons.
The Genetic Code
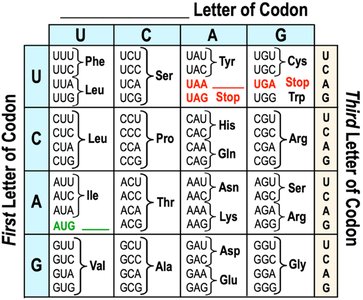
Decoding the Genetic Code
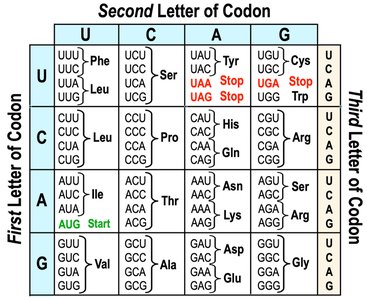
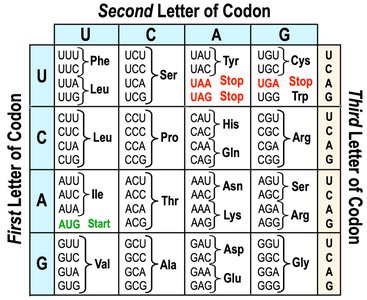
The genetic code is a set of rules by which the sequence of nucleotides in mRNA is translated into the sequence of amino acids in a protein. It is read in triplets called codons, each specifying a particular amino acid or a stop signal.
Redundancy: Multiple codons can code for the same amino acid.
Start Codon: AUG (codes for Methionine) signals the start of translation.
Stop Codons: UAA, UAG, UGA signal the end of translation.
Translation: From RNA to Protein
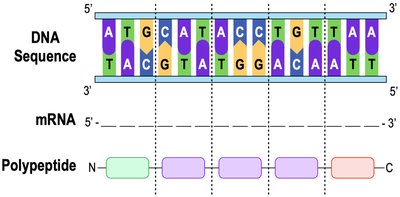
Introduction to Translation
Translation is the process by which ribosomes synthesize proteins using the sequence of codons in mRNA. This process involves the coordinated action of mRNA, tRNA, and rRNA.
Ribosomes: Complexes of rRNA and protein that facilitate the translation of mRNA into polypeptides.
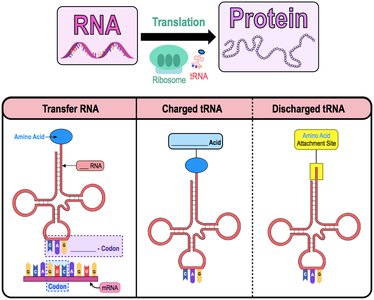
tRNA: Delivers specific amino acids to the ribosome by matching its anticodon to the mRNA codon.
Charged tRNA: tRNA attached to its corresponding amino acid.
Discharged tRNA: tRNA that has released its amino acid.
Ribosome Structure and tRNA Binding Sites
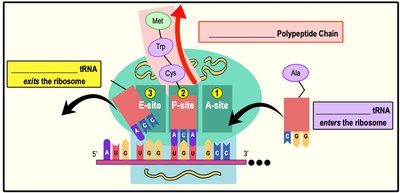
Ribosomes are composed of two subunits (large and small) and have three binding sites for tRNA:
A (Aminoacyl) Site: Holds the tRNA carrying the next amino acid.
P (Peptidyl) Site: Holds the tRNA with the growing polypeptide chain.
E (Exit) Site: Where discharged tRNAs leave the ribosome.
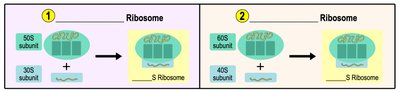
Prokaryotic ribosomes are 70S (50S + 30S), while eukaryotic ribosomes are 80S (60S + 40S).
Steps of Translation
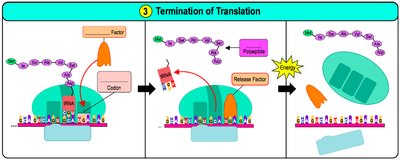
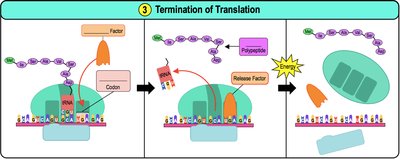
Translation occurs in three main steps: initiation, elongation, and termination.
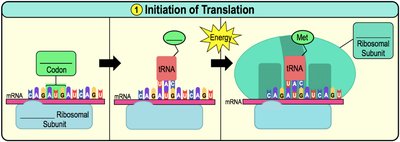
Initiation: The small ribosomal subunit binds to mRNA and the initiator tRNA (carrying methionine) binds to the start codon (AUG). The large subunit then joins.
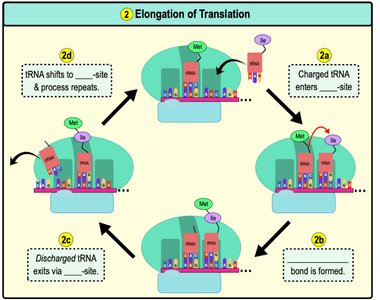
Elongation: Amino acids are added one by one to the growing polypeptide chain. The ribosome moves along the mRNA in the 5' to 3' direction.
Termination: When a stop codon is reached, release factors bind, causing the polypeptide to be released and the ribosome to disassemble.
Post-Translational Modifications
Protein Modifications After Translation
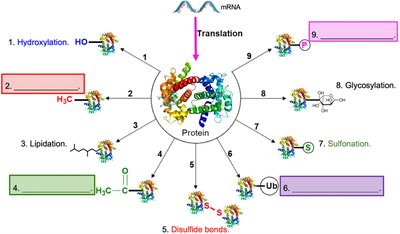
After translation, proteins often undergo post-translational modifications (PTMs) that regulate their activity, stability, and function. Common PTMs include:
Methylation
Acetylation
Ubiquitination
Phosphorylation
Glycosylation: Addition of carbohydrates to proteins.
Comparison: Transcription vs. Translation
Key Differences and Similarities
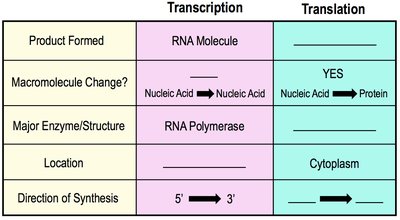
Transcription | Translation | |
|---|---|---|
Product Formed | RNA Molecule | Polypeptide (Protein) |
Macromolecule Change? | Nucleic Acid → Nucleic Acid | Nucleic Acid → Protein |
Major Enzyme/Structure | RNA Polymerase | Ribosome |
Location | Nucleus (Eukaryotes) | Cytoplasm |
Direction of Synthesis | 5' → 3' | N-terminus → C-terminus |
Mutations
Types and Effects of Mutations
Mutations are permanent changes in the DNA sequence. They can affect gene expression and protein function, and may be harmful, beneficial, or neutral. Mutations can occur spontaneously or be induced by environmental factors (mutagens).
Point Mutation: Change in a single nucleotide (can be silent, missense, or nonsense).
Frameshift Mutation: Insertion or deletion of nucleotides that alters the reading frame.
Silent Mutation: Does not change the amino acid sequence.
Missense Mutation: Changes one amino acid in the protein.
Nonsense Mutation: Introduces a premature stop codon, truncating the protein.
*Additional info: Mutations are a major source of genetic diversity and evolution.*
